Teodors Pantelejevs, Pedro Zuazua-Villar, Oliwia Koczy, Stephen J Walsh, Naomi Stephanie Robertson, Andrew J. Counsell, David R Spring, Jessica A. Downs and Marko Hyvönen
Chemical Science, accepted manuscript (2023)
DOI: 10.1039/D3SC03331G
PDB coordinates: 8c3j (3D view), 8br9 (3D view), 8c3n (3D view)

Abstract
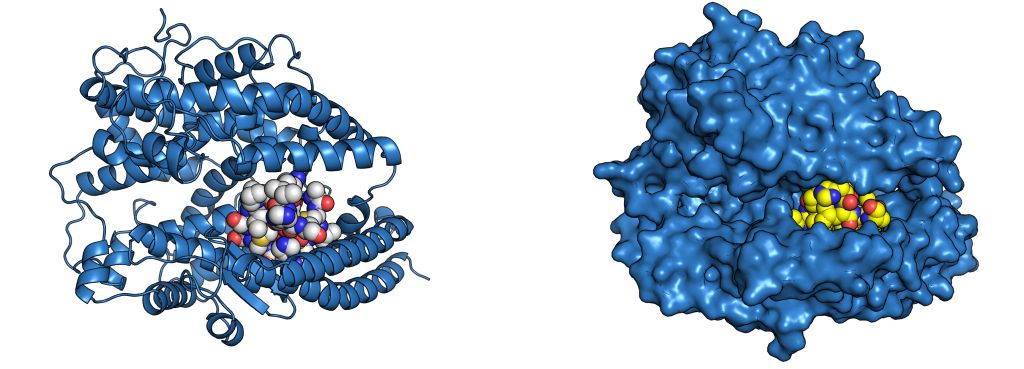
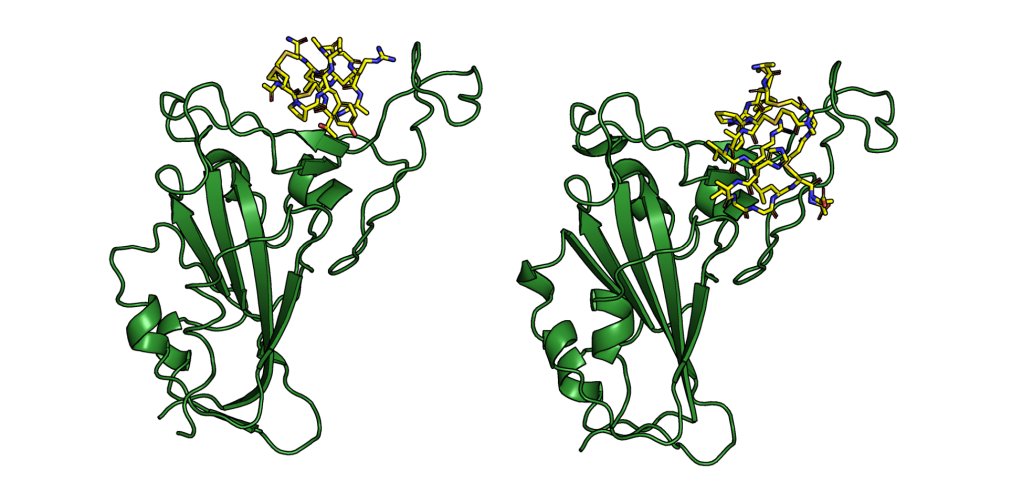
Stapling is a macrocyclisation method that connects amino acid side chains of a peptide to improve its pharmacological properties. We describe an approach for stapled peptide preparation and biochemical evaluation that combines recombinant expression of fusion constructs of target peptides and cysteine-reactive divinyl-heteroaryl chemistry as an alternative to solid-phase synthesis. We then employ this workflow to prepare and evaluate BRC-repeat-derived inhibitors of the RAD51 recombinase, showing that a diverse range of secondary structure elements in the BRC repeat can be stapled without compromising binding and function. Using X-ray crystallography, we elucidate the atomic-level features of the staple moieties. We then demonstrate that BRC-repeat-derived stapled peptides can disrupt RAD51 function in cells following ionising radiation treatment.